Databases & Servers
I have been actively involved as a developer and co-developer in several scientific databases and web servers that support computational biology, peptide design, and drug discovery research.

pLM4CPPs — AI/ML Predictor for CPPs
Deep learning architecture combining protein language models and convolutional neural networks for high-accuracy prediction of cell-penetrating peptides.
Journal of Chemical Information and Modeling (2025) — https://doi.org/10.1021/acs.jcim.4c01338
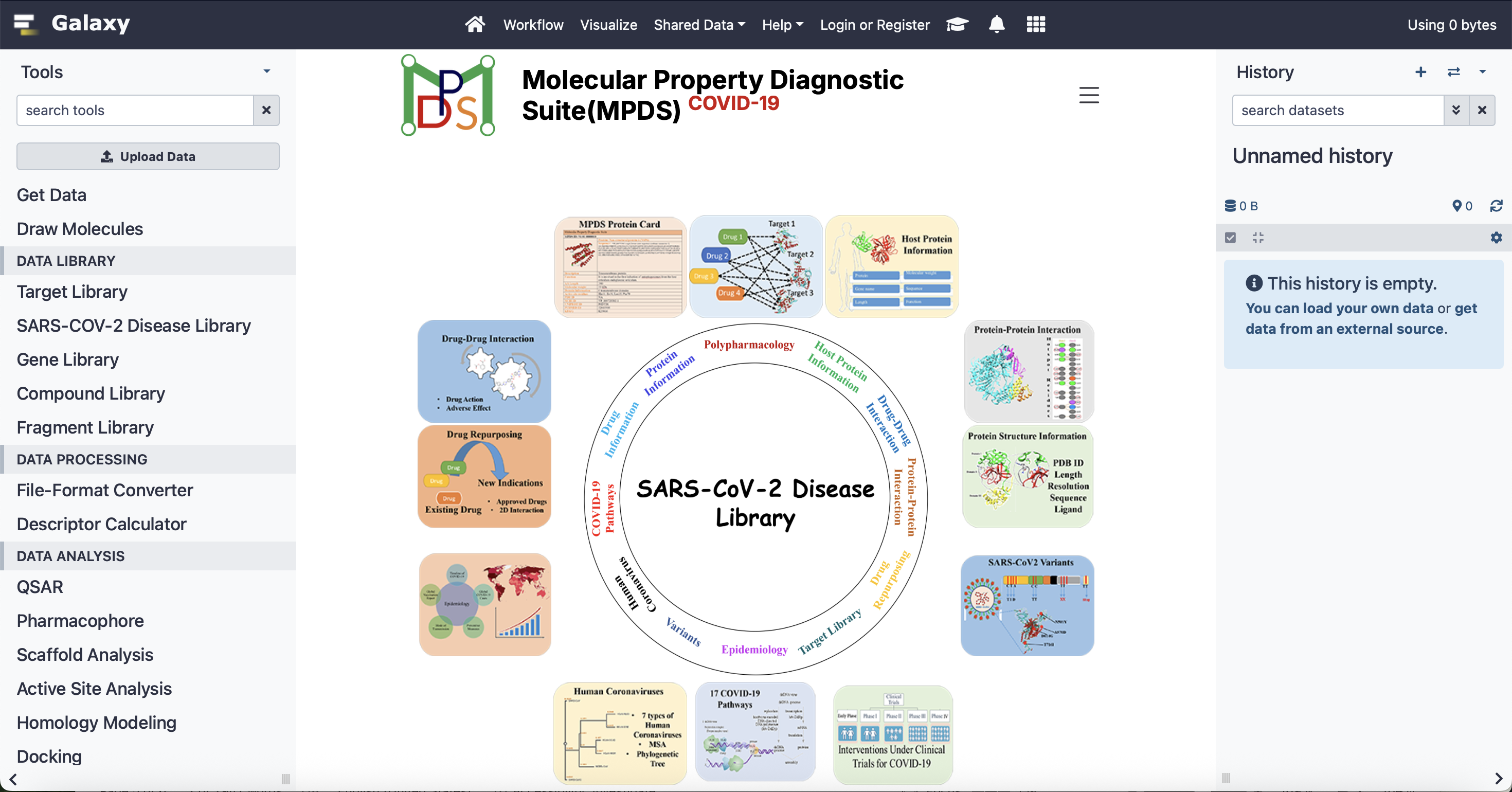
MPDS COVID-19 Web Portal
An open-source drug discovery portal integrating genes, proteins, pathways, binding sites, and AI-powered compound analysis with a database of over 149 million molecules and 107,000 fragments.
GigaByte (2024) — https://doi.org/10.1101/2023.08.29.555437
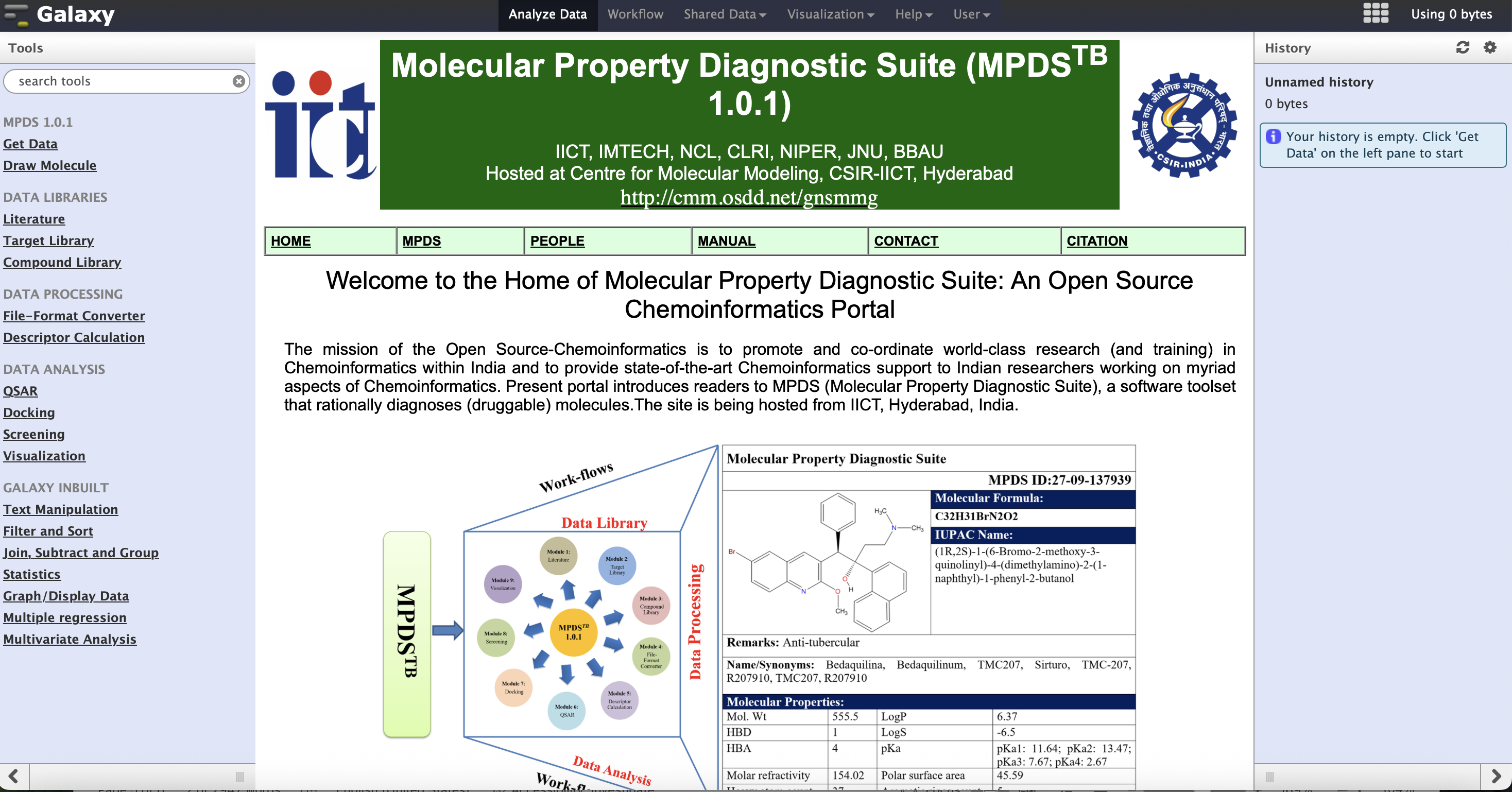

MPDS TB Web Portal
An integrated computational suite for tuberculosis research, providing cheminformatics workflows, CADD tools, and unique MPDS identifiers for compound tracking.
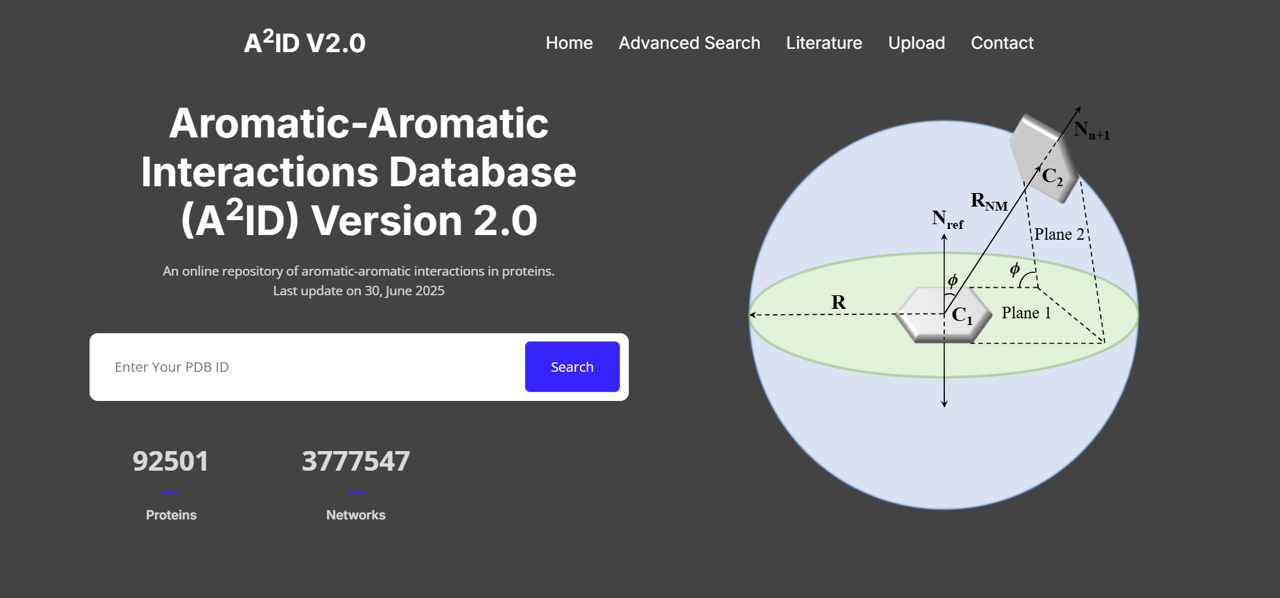
Aromatic–Aromatic Interactions Database (A2ID)
A curated database of π–π interaction motifs involving Trp, Tyr, Phe, and His residues, supporting analysis of aromatic stabilization in protein structures.
International Journal of Biological Macromolecules (2023) — https://doi.org/10.1016/j.ijbiomac.2023.127207
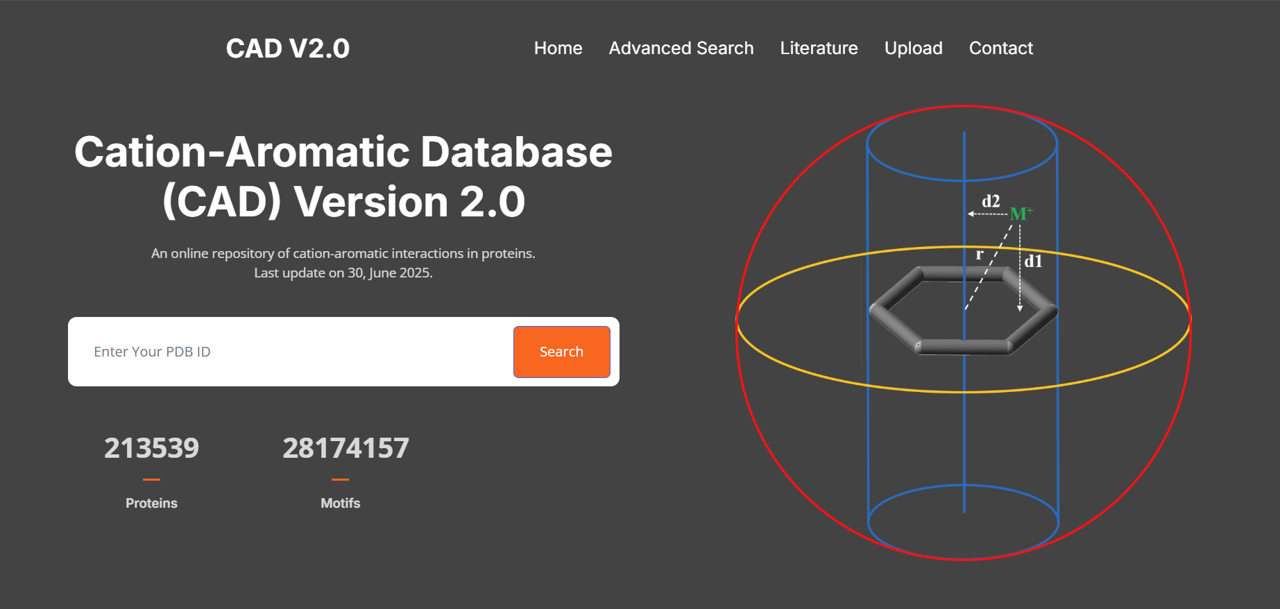
Cation–Aromatic Database (CAD 2.0)
A comprehensive repository of over 27 million cation–aromatic interaction motifs, including geometric parameters, residue preferences, and classification of interaction types.
Proteins (2023) — https://doi.org/10.1002/prot.26600